Medical Imaging
Multivariate semi-blind deconvolution of fMRI time series
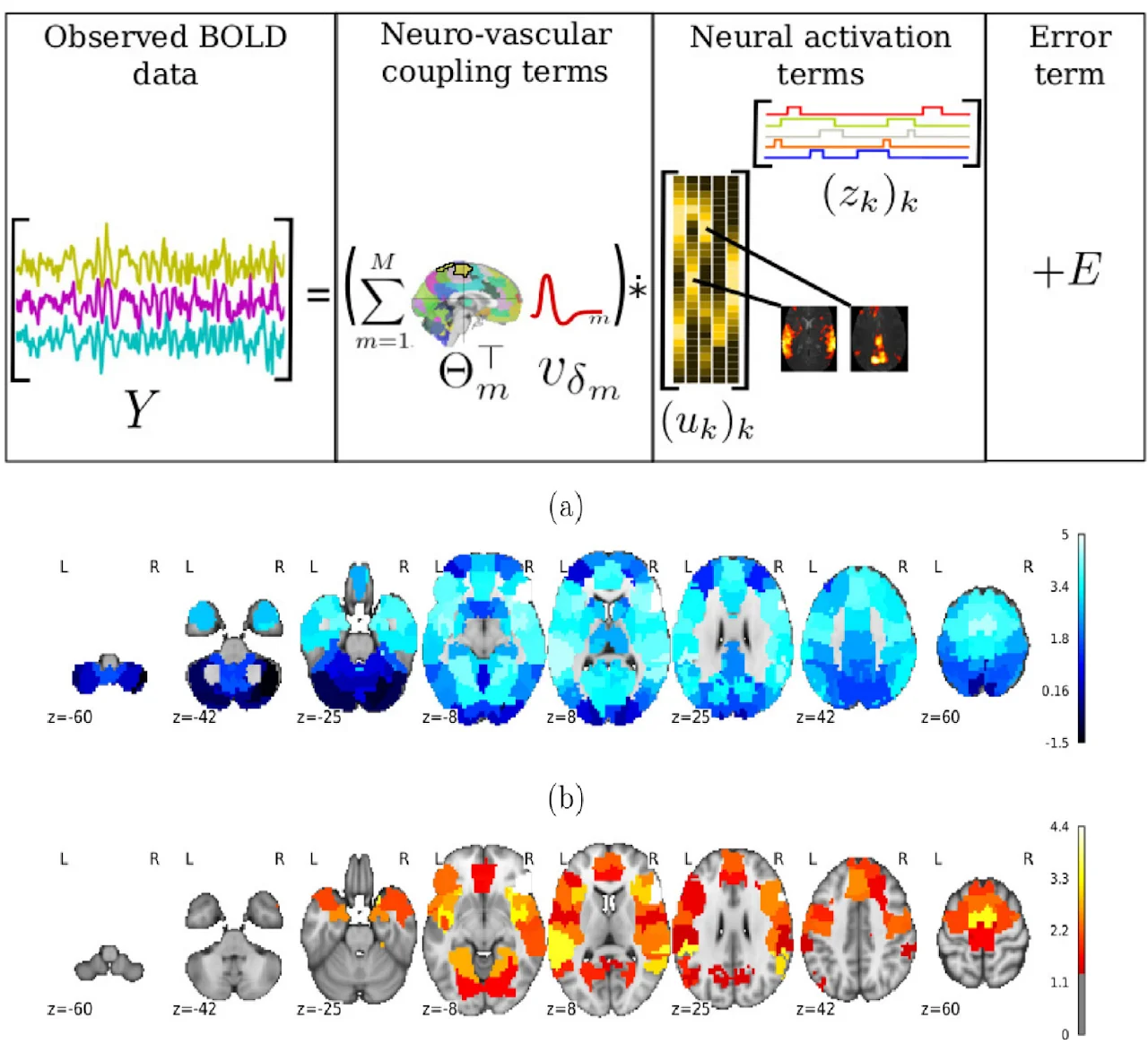
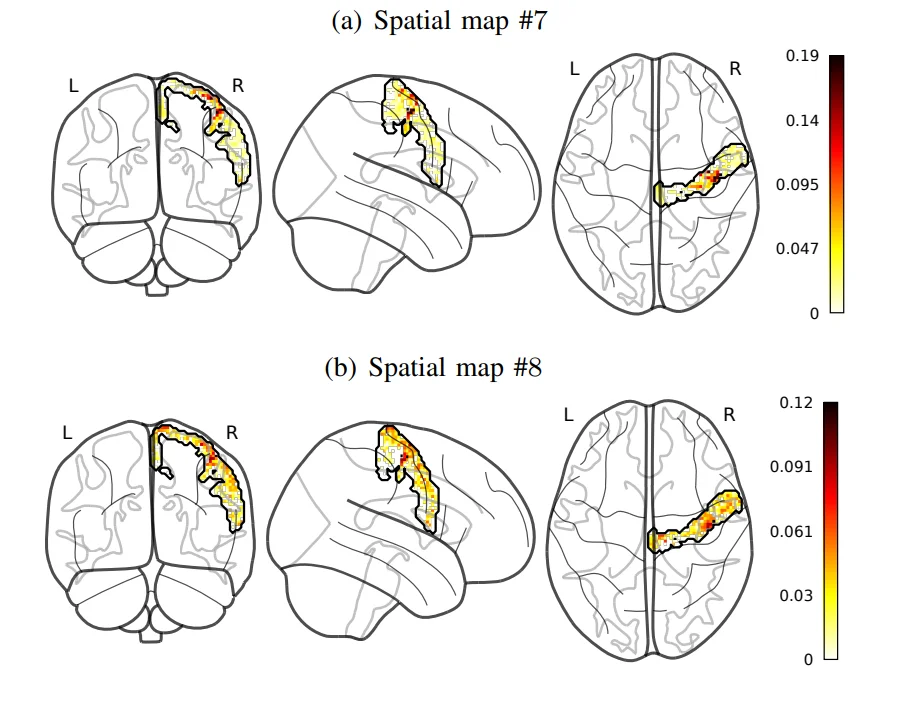
Whole brain estimation of the haemodynamic response function (HRF) in functional magnetic resonance imaging (fMRI) is critical for understanding neurovascular coupling. Traditional approaches rely on task-fMRI data, rendering them inoperative on resting-state fMRI (rsfMRI). To address this, we formulate the joint estimation of HRF shapes and spatio-temporal neural representations as a multivariate semi-blind deconvolution problem in a paradigm-free setting. We express neural activity signals using piecewise constant temporal atoms paired with sparse spatial maps, while introducing a haemodynamic parcellation governed by temporal dilations. Validated on synthetic and real rsfMRI data, our fast alternating minimization algorithm efficiently discriminates health conditions at the population level. Applied to the UK Biobank dataset, we statistically demonstrate that pathologies like stroke or normal brain aging induce measurable haemodynamic delays in specific brain areas, serving as an effective predictive feature.
fMRI BOLD signal decomposition using a multivariate low-rank model
Standard fMRI data analysis typically relies on voxelwise linear modeling with maximum likelihood estimation, whereas resting-state data often require multivariate approaches like PCA or ICA to extract information in a data-driven manner. We propose a novel low-rank model for fMRI BOLD data that shifts away from standard spatial dimension reduction. Instead, our model utilizes convolutional sparse coding between the hemodynamic system and specific temporal atoms that code for neural activity. A rank-1 constraint is applied to each temporal atom to accurately map its spatial influence across the brain. Formulated as a nonconvex optimization problem within a variational framework, we use an efficient alternate minimization algorithm to jointly estimate neural signals and spatial maps. Validation demonstrates that our approach uncovers competitive neural fingerprints while offering a substantially richer decomposition in both time and space.
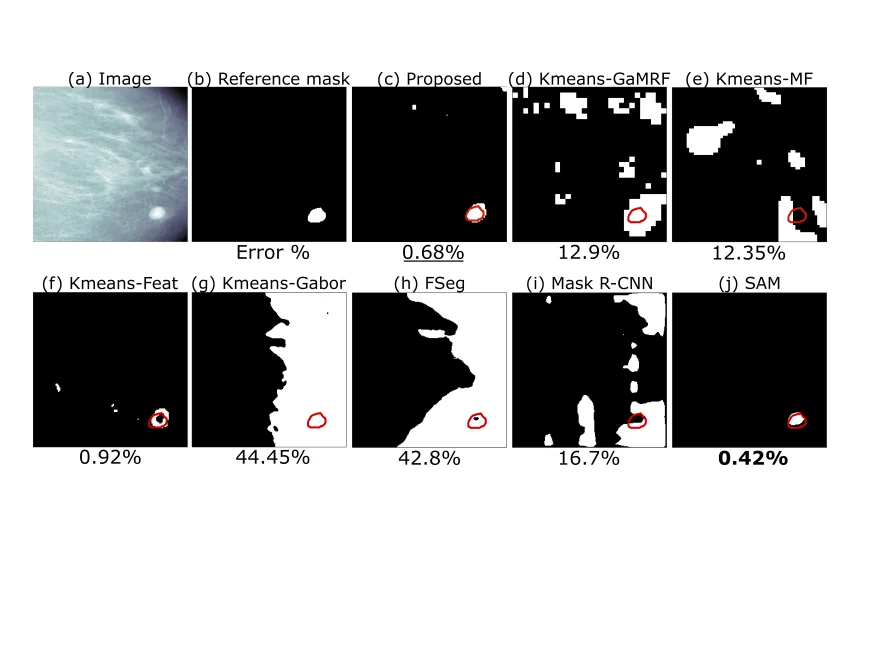
Bayesian Multifractal Image Segmentation
Multifractal analysis (MFA) effectively characterizes image textures by describing spatial fluctuations in local regularity. However, natural images typically contain multiple complex textures and their associated multifractal properties. We introduce an unsupervised Bayesian multifractal segmentation method that jointly models and segments multifractal textures at the pixel-level. This approach develops a computationally efficient parameter estimation model for wavelet leaders, assigning specific multifractality parameters to different image regions. Furthermore, we leverage a multiscale Potts Markov random field as a prior to capture the inherent spatial and cross-scale correlations between wavelet leader labels. Utilizing a Gibbs sampling methodology, we draw samples from the posterior distribution of the unknown model parameters. Numerical evaluations demonstrate that our Bayesian approach achieves superior performance and effectiveness in multifractal image segmentation when compared to both traditional unsupervised techniques and modern deep learning models.
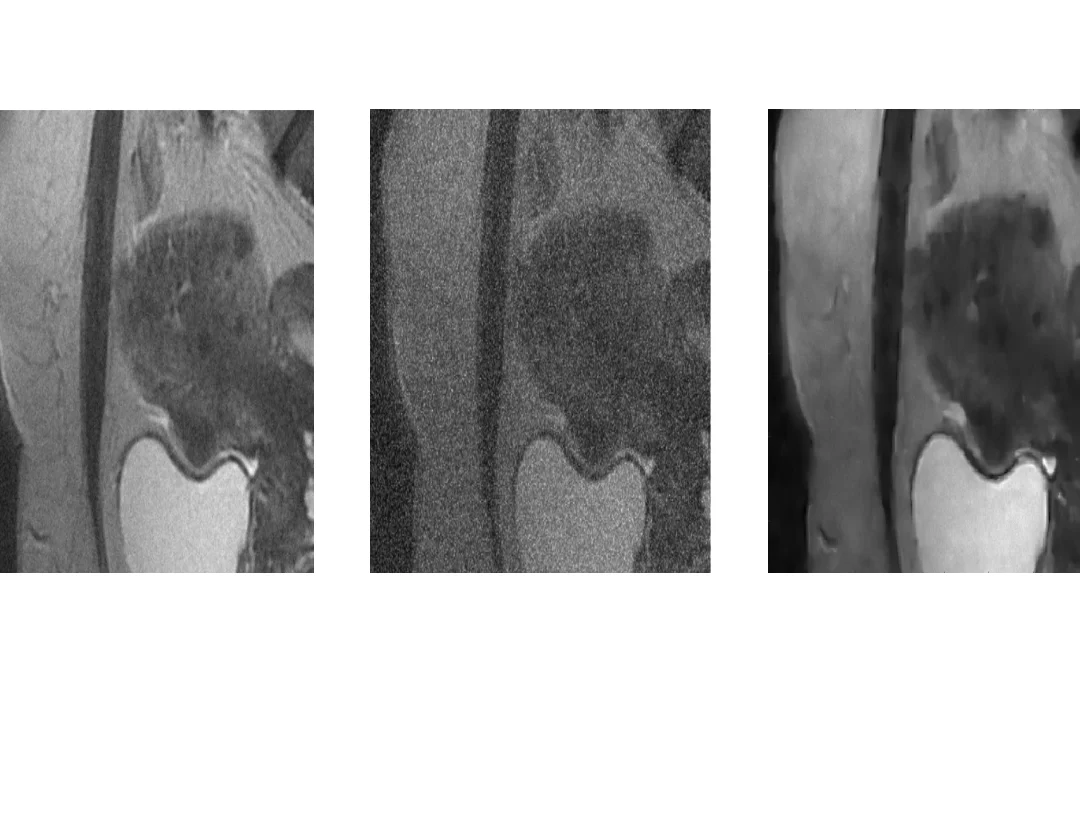
Fusion of Magnetic Resonance and Ultrasound Images Using Guided Filtering: Application to Endometriosis Surgery
This paper studies a new fusion method designed for magnetic resonance (MR) and ultrasound (US) images, with a specific focus on endometriosis diagnosis. The proposed method is based on guided filtering, leveraging the advantages of this technique to enhance the quality of fused images. The fused image is a weighted average of base and detail images from the MR and US images. The weights assigned to the US image account for the presence of speckle noise, a common challenge in US imaging whereas the weights assigned to the MR image allow the contrast of the fused image to be enhanced. The effectiveness of the method is evaluated using synthetic and phantom data, showing promising results. The image provided by the proposed fusion method holds potential for enhancing visualization and aiding decision-making in endometriosis surgery, offering a valuable contribution to the field of medical image fusion.